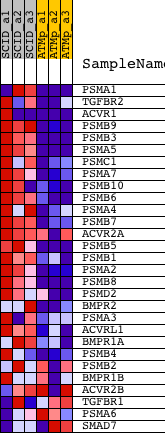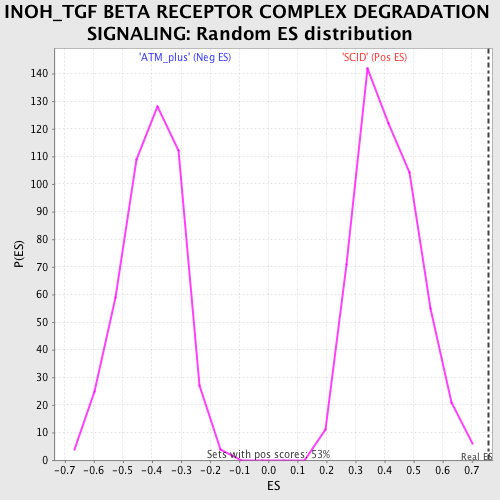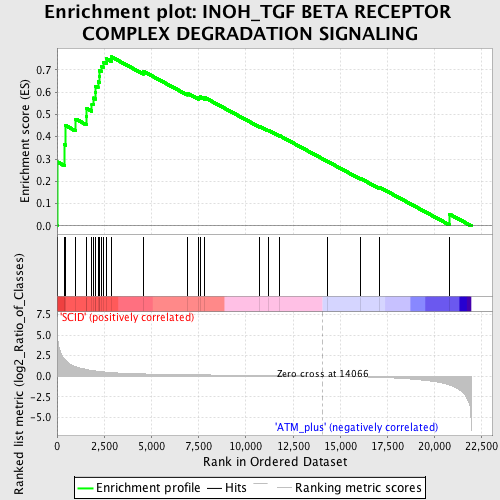
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_01_SCID_versus_ATM_plus.phenotype_SCID_versus_ATM_plus.cls #SCID_versus_ATM_plus.phenotype_SCID_versus_ATM_plus.cls #SCID_versus_ATM_plus_repos |
| Phenotype | phenotype_SCID_versus_ATM_plus.cls#SCID_versus_ATM_plus_repos |
| Upregulated in class | SCID |
| GeneSet | INOH_TGF BETA RECEPTOR COMPLEX DEGRADATION SIGNALING |
| Enrichment Score (ES) | 0.7581873 |
| Normalized Enrichment Score (NES) | 1.8435116 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.006431661 |
| FWER p-Value | 0.094 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PSMA1 | 1415695_at 1436769_at 1436770_x_at 1458292_at | 8 | 6.245 | 0.2868 | Yes | ||
| 2 | TGFBR2 | 1422019_at 1425444_a_at 1426397_at 1443115_at 1458607_at | 393 | 2.063 | 0.3641 | Yes | ||
| 3 | ACVR1 | 1416786_at 1416787_at 1448460_at 1457551_at | 455 | 1.922 | 0.4497 | Yes | ||
| 4 | PSMB9 | 1450696_at | 978 | 1.164 | 0.4794 | Yes | ||
| 5 | PSMB3 | 1417052_at | 1556 | 0.815 | 0.4905 | Yes | ||
| 6 | PSMA5 | 1424681_a_at 1434356_a_at | 1570 | 0.805 | 0.5269 | Yes | ||
| 7 | PSMC1 | 1416005_at | 1843 | 0.691 | 0.5463 | Yes | ||
| 8 | PSMA7 | 1423567_a_at 1423568_at | 1912 | 0.664 | 0.5737 | Yes | ||
| 9 | PSMB10 | 1448632_at | 2021 | 0.629 | 0.5977 | Yes | ||
| 10 | PSMB6 | 1448822_at | 2044 | 0.622 | 0.6253 | Yes | ||
| 11 | PSMA4 | 1460339_at | 2170 | 0.587 | 0.6465 | Yes | ||
| 12 | PSMB7 | 1416240_at | 2230 | 0.572 | 0.6702 | Yes | ||
| 13 | ACVR2A | 1437382_at 1439643_at 1445026_at 1451004_at | 2241 | 0.570 | 0.6959 | Yes | ||
| 14 | PSMB5 | 1415676_a_at | 2348 | 0.542 | 0.7160 | Yes | ||
| 15 | PSMB1 | 1420052_x_at 1420053_at 1448166_a_at | 2470 | 0.514 | 0.7341 | Yes | ||
| 16 | PSMA2 | 1448206_at | 2629 | 0.483 | 0.7491 | Yes | ||
| 17 | PSMB8 | 1422962_a_at 1444619_x_at | 2873 | 0.439 | 0.7582 | Yes | ||
| 18 | PSMD2 | 1415831_at | 4564 | 0.273 | 0.6936 | No | ||
| 19 | BMPR2 | 1419616_at 1434310_at 1441652_at | 6900 | 0.181 | 0.5953 | No | ||
| 20 | PSMA3 | 1448442_a_at | 7498 | 0.164 | 0.5756 | No | ||
| 21 | ACVRL1 | 1435825_at 1451604_a_at | 7592 | 0.161 | 0.5788 | No | ||
| 22 | BMPR1A | 1425491_at 1425492_at 1425493_at 1425494_s_at 1445413_at 1451729_at | 7823 | 0.154 | 0.5753 | No | ||
| 23 | PSMB4 | 1438984_x_at 1451205_at | 10735 | 0.083 | 0.4463 | No | ||
| 24 | PSMB2 | 1444377_at 1448262_at 1459641_at | 11183 | 0.073 | 0.4292 | No | ||
| 25 | BMPR1B | 1422872_at 1437312_at 1443720_s_at | 11779 | 0.058 | 0.4047 | No | ||
| 26 | ACVR2B | 1419140_at 1439856_at | 14340 | -0.008 | 0.2882 | No | ||
| 27 | TGFBR1 | 1420893_a_at 1420894_at 1420895_at 1446946_at | 16054 | -0.073 | 0.2134 | No | ||
| 28 | PSMA6 | 1416506_at 1435316_at 1435317_x_at 1437144_x_at | 17094 | -0.142 | 0.1725 | No | ||
| 29 | SMAD7 | 1423389_at 1440952_at 1443771_x_at | 20786 | -1.054 | 0.0525 | No |